Publications
2025
-
 ModalTune: Fine-Tuning Slide-Level Foundation Models with Multi-Modal Information for Multi-task Learning in Digital PathologyVishwesh Ramanathan*, Tony Xu*, Pushpak Pati, and 3 more authorsInternational Conference on Computer Vision (ICCV), 2025
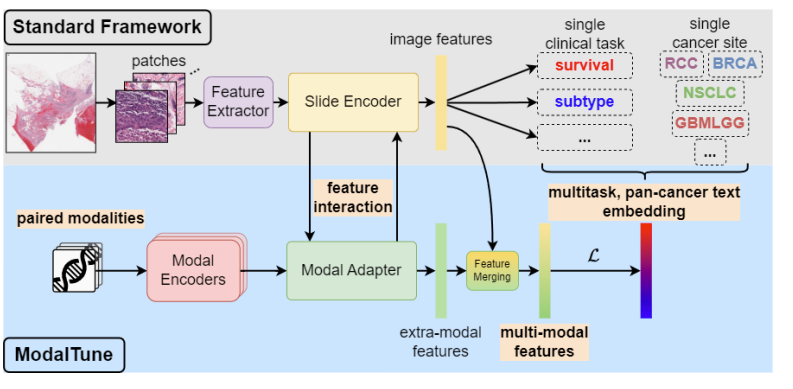
ModalTune: Fine-Tuning Slide-Level Foundation Models with Multi-Modal Information for Multi-task Learning in Digital PathologyVishwesh Ramanathan*, Tony Xu*, Pushpak Pati, and 3 more authorsInternational Conference on Computer Vision (ICCV), 2025Prediction tasks in digital pathology are challenging due to the massive size of whole-slide images (WSIs) and the weak nature of training signals. Advances in computing, data availability, and self-supervised learning (SSL) have paved the way for slide-level foundation models (SLFMs) that can improve prediction tasks in low-data regimes. However, working with these models is challenging, with issues such as catastrophic forgetting during fine-tuning and underutilization of shared information between tasks and modalities. To overcome these two challenges, we propose ModalTune, a novel fine-tuning framework which introduces the Modal Adapter to integrate new modalities without modifying SLFM weights. Additionally, we use large-language models (LLMs) to encode labels as text, capturing semantic relationships and enhancing generalization across multiple tasks and cancer types in a single training recipe. ModalTune achieves state-of-the-art (SOTA) results against both uni-modal and multi-modal models across four cancer types, jointly improving survival and cancer subtype prediction while remaining competitive in pan-cancer settings. Additionally, we show ModalTune is highly generalizable to two out-of-distribution (OOD) datasets. To our knowledge, this is the first unified fine-tuning framework for multi-modal, multi-task, and pan-cancer modeling in digital pathology.
-
 Tumor-infiltrating lymphocytes in breast cancer through artificial intelligence: biomarker analysis from the results of the TIGER challengeMart Rijthoven, Witali Aswolinskiy, Leslie Tessier, and 8 more authorsmedRxiv, 2025
Tumor-infiltrating lymphocytes in breast cancer through artificial intelligence: biomarker analysis from the results of the TIGER challengeMart Rijthoven, Witali Aswolinskiy, Leslie Tessier, and 8 more authorsmedRxiv, 2025The prognostic significance of tumor-infiltrating lymphocytes (TILs) in breast cancer has been recognized for over a decade. Although histology-based scoring recommendations exist to standardize visual TILs assessment, interobserver agreement and reproducibility are hampered by heterogeneous infiltration patterns, highlighting the importance of computational approaches. Despite advances to automate TILs quantification, adoption of computational models has been hindered by lack of consensus on scoring methods and lack of large-scale benchmarks. To address these limitations, we launched the international TIGER challenge, a public competition to build open-source computational TILs (cTILs) models in digital pathology. Here, we present the largest comprehensive multi-centric validation of multiple cTILs methods on surgical resections and biopsies using 3,708 Triple Negative Breast Cancer (TNBC) and human epidermal growth factor receptor 2 positive (HER2+) breast cancers from clinical practice and phase 3 clinical trials. We report benchmarks on image analysis performance of each method and show the strong agreement of cTILs with panels of pathologists. We show the positive association of cTILS with response after neoadjuvant therapy in HER2-positive, superior to visually scored TILs. We also show that cTILs add independent information to clinical variables in surgically resected TNBC but not in HER2-positive disease and breast biopsies
2024
- Med-ImageTools: An open-source Python package for robust data processing pipelines and curating medical imaging dataSejin Kim, Michal Kazmierski, Kevin Qu, and 7 more authorsF1000Research, 2024
Machine learning and AI promise to revolutionize the way we leverage medical imaging data for improving care but require large datasets to train computational models that can be implemented in clinical practice. However, processing large and complex medical imaging datasets remains an open challenge. To address this issue, we developed Med-ImageTools, a new Python open-source software package to automate data curation and processing while allowing researchers to share their data processing configurations more easily, lowering the barrier for other researchers to reproduce published works. We have demonstrated the efficiency of Med-ImageTools across three different datasets, resulting in significantly reduced processing times. The AutoPipeline feature will improve the accessibility of raw clinical datasets on public archives, such as the Cancer Imaging Archive (TCIA), the largest public repository of cancer imaging, allowing machine learning researchers to process analysis-ready formats without requiring deep domain knowledge
-
 Ensemble of Prior-guided Expert Graph Models for Survival Prediction in Digital PathologyVishwesh Ramanathan*, Pushpak Pati*, Matthew McNeil, and 1 more authorIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2024
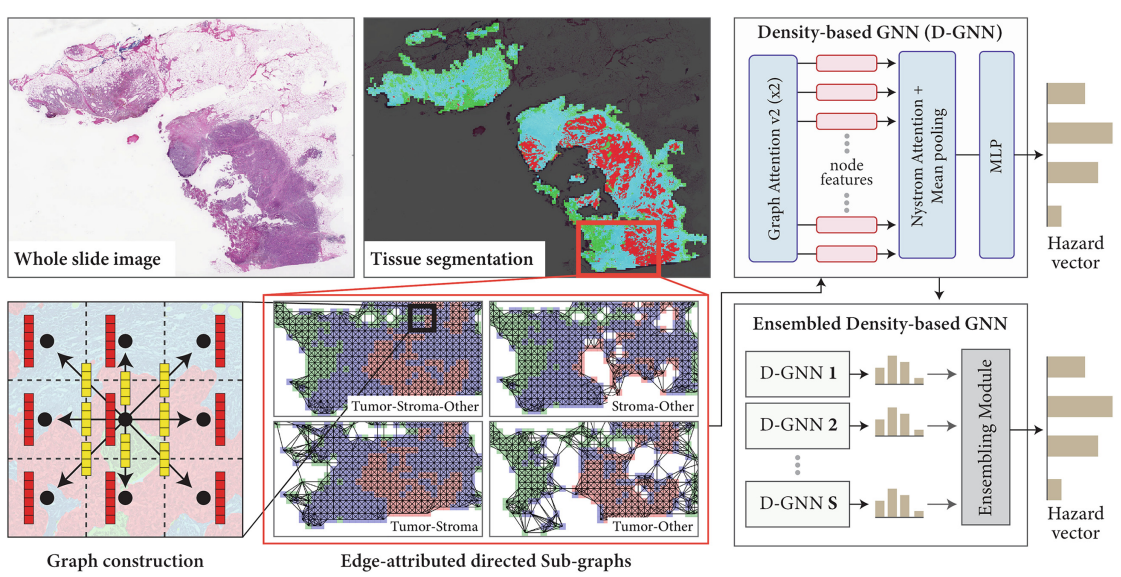
Ensemble of Prior-guided Expert Graph Models for Survival Prediction in Digital PathologyVishwesh Ramanathan*, Pushpak Pati*, Matthew McNeil, and 1 more authorIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2024Survival prediction in pathology is a dynamic research field focused on identifying predictive biomarkers to enhance cancer survival models, providing valuable guidance for clinicians in treatment decisions. Graph-based methods, especially Graph Neural Networks (GNNs) leveraging rich interactions among different biological entities, have recently successfully predicted survival. However, the inherent heterogeneity among the entities within tissue slides significantly challenges the learning of GNNs. GNNs, operating with the homophily assumption, diffuse the intricate interactions among heterogeneous tissue entities in a tissue microenvironment. Further, the convoluted downstream task relevant information is not effectively exploited by graph-based methods when working with large slide-graphs. We propose a novel prior-guided, edge-attributed tissue-graph construction to address these challenges, followed by an ensemble of expert graph-attention survival models. Our method exploits diverse prognostic factors within numerous targeted tissue subgraphs of heterogeneous large slide-graphs. Our method achieves state-of-the-art results on four cancer types, improving overall survival prediction by 4.33% compared to the competing methods
-
 Detecting Noisy Labels with Repeated Cross-ValidationsJianan Chen*, Vishwesh Ramanathan*, Tony Xu, and 1 more authorIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2024
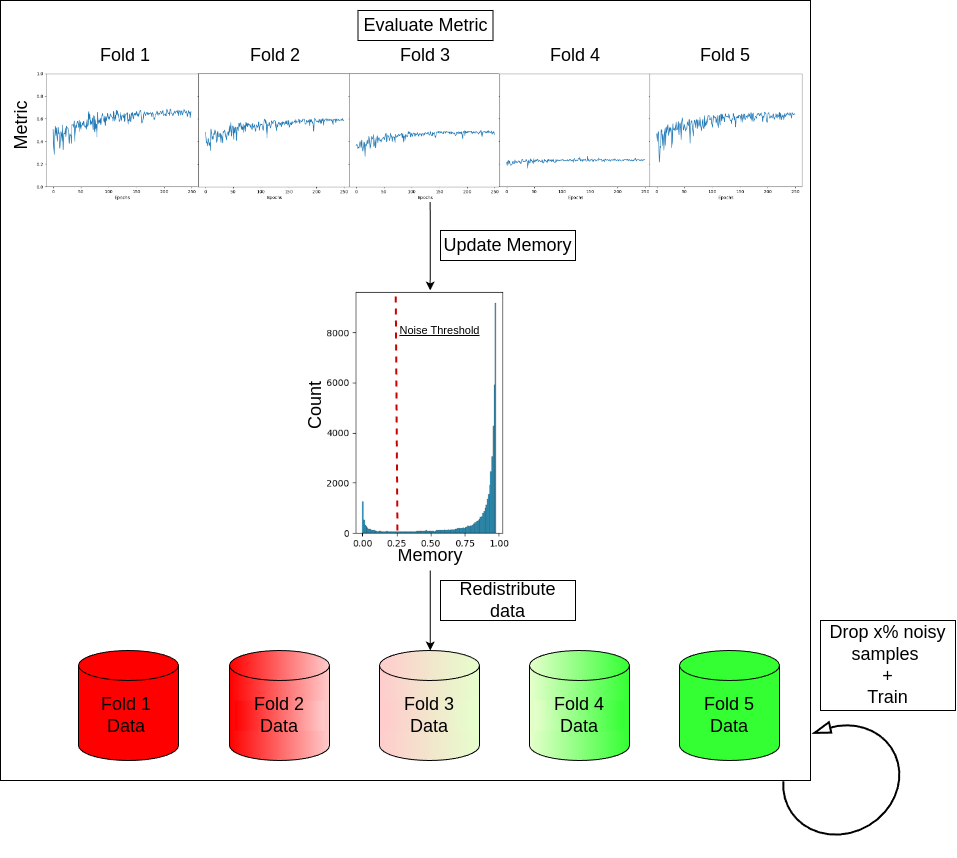
Detecting Noisy Labels with Repeated Cross-ValidationsJianan Chen*, Vishwesh Ramanathan*, Tony Xu, and 1 more authorIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2024Machine learning models experience deteriorated performance when trained in the presence of noisy labels. This is particularly problematic for medical tasks, such as survival prediction, which typically face high label noise complexity with few clear-cut solutions. Inspired by the large fluctuations across folds in the cross-validation performance of survival analyses, we design Monte-Carlo experiments to show that such fluctuation could be caused by label noise. We propose two novel and straightforward label noise detection algorithms that effectively identify noisy examples by pinpointing the samples that more frequently contribute to inferior cross-validation results. We first introduce Repeated Cross-Validation (ReCoV), a parameter-free label noise detection algorithm that is robust to model choice. We further develop fastReCoV, a less robust but more tractable and efficient variant of ReCoV suitable for deep learning applications. Through extensive experiments, we show that ReCoV and fastReCoV achieve state-of-the-art label noise detection performance in a wide range of modalities, models and tasks, including survival analysis, which has yet to be addressed in the literature
- Spatial analysis of immune cells in breast cancer using k-nearest neighbor graphs and Louvain-community clustering of immunofluorescent protein multiplexing imagesAlison M Cheung, Wenchao Han, Snow Zhou, and 5 more authorsIn 17th International Workshop on Breast Imaging (IWBI 2024), 2024
Immune phenotype data, specifically the description of densities and spatial distribution of immune cells are now frequently included in the clinical pathology report as these features of the cells in the tumor microenvironment (TME) have shown to be associated with prognosis. In addition, immune-therapeutics, which aim at manipulating the patients’ immune system to kill cancer cells, have recently been approved for treatment of triple-negative breast cancers (TNBCs). Thus, quantifying the immune phenotype of the cancer could be important both for prognostication, and for prediction of therapy response. We have studied the immune phenotype of 42 breast cancers using immunofluorescence protein multiplexing and quantitative image analysis. After sectioning, formalin-fixed paraffin-embedded tissues were sequentially stained with a panel of fluorescently-labelled antibodies and imaged with the multiplexer (Cell DIVE, Leica Biosystems). Composite images of antibody-stained sections were then analysed using specialized digital pathology software (HALO, Indica Labs). Binary thresholding was conducted to identify and quantify densities of various immune lineage subsets (T lymphocytes and macrophages). Their cellular localisation was mapped and the spatial features of cellular arrangement were evaluated using a k-nearest neighbor graph (KNNG) method and Louvain community-proximity clustering. The spatial relationship of various immune and cancer cell types was quantified to assess whether cellular arrangements and structures differed among breast cancer subtypes. Our work demonstrates the use of molecular and cellular imaging in quantifying features of the tumor microenvironment in breast cancer classification, and the application of KNNG in studying spatial biology
2023
- Identification of molecular cell type of breast cancer on digital histopathology images using deep learning and multiplexed fluorescence imagingWenchao Han, Alison M Cheung, Vishwesh Ramanathan, and 4 more authorsIn Medical Imaging 2023: Digital and Computational Pathology, 2023
ER, PR (estrogen, progesterone receptor), and HER2 (human epidermal growth factor receptor 2) status are assessed using immunohistochemistry and reported in standard clinical workflows as they provide valuable information to help treatment planning. The protein Ki67 has also been suggested as a prognostic biomarker but is not routinely evaluated clinically due to insufficient quality assurance. The routine pathological practice usually relies on small biopsies, such that the reduction in consumption is necessary to save materials for special assays. For this purpose, we developed and validated an automatic system for segmenting and identifying the (ER, PR, HER2, Ki67) positive cells from hæmatoxylin and eosin (H&E) stained tissue sections using multiplexed immunofluorescence (MxIF) images at cellular level as a reference standard. In this study, we used 100 tissue-microarray cores sampled from 56 cases of invasive breast cancer. For ER, we extracted cell nucleus images (HoverNet) from the H&E images and assigned each cell nucleus as ER positive vs. negative based on the corresponding MxIF signals (whole cell segmentation with DeepCSeg) upon H&E to MxIF image registration. We trained a Res-Net 18 and validated the model on a separate test-set for classifying the cells as positive vs. negative for ER, and performed the same experiment for the other three markers. We obtained area-under-the- receiver-operating-characteristic-curves (AUCs) of 0.82 (ER), 0.85 (PR), 0.75 (HER2), 0.82 (Ki67) respectively. Our study demonstrates the feasibility of using machine learning to identify molecular status at cellular level directly from the H&E slides
-
 Ink removal in whole slide images using hallucinated dataVishwesh Ramanathan, Wenchao Han, Dina Bassiouny, and 2 more authorsIn Medical Imaging 2023: Digital and Computational Pathology, 2023
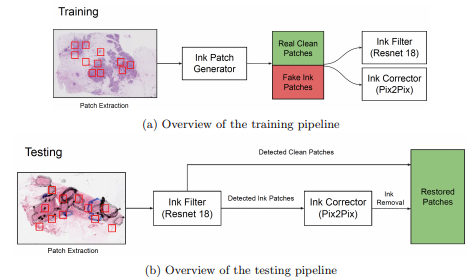
Ink removal in whole slide images using hallucinated dataVishwesh Ramanathan, Wenchao Han, Dina Bassiouny, and 2 more authorsIn Medical Imaging 2023: Digital and Computational Pathology, 2023Pathologists regularly use ink markings on histopathology slides to highlight specific areas of interest or orientation, making it an integral part of the workflow. Unfortunately, digitization of these ink-annotated slides hinders any computer-aided analyses, particularly deep learning algorithms, which require clean data free from artifacts. We propose a methodology that can identify and remove the ink markings for the purpose of computational analyses. We propose a two-stage network with a binary classifier for ink filtering and Pix2Pix for ink removal. We trained our network by artificially generating pseudo ink markings using only clean slides, requiring no manual annotation or curation of data. Furthermore, we demonstrate our algorithm’s efficacy over an independent dataset of H&E stained breast carcinoma slides scanned before and after the removal of pen markings. Our quantitative analysis shows promising results, achieving 98.7% accuracy for the binary classifier. For Pix2Pix, we observed a 65.6% increase in structure similarity index, a 21.3% increase in peak signal-to-noise ratio, and a 30% increase in visual information fidelity. As only clean slides are required for training, the pipeline can be adapted to multiple colors of ink markings or new domains, making it easy to deploy over different sets of histopathology slides
-
 Self Supervised Multi-view Graph Representation Learning in Digital PathologyVishwesh Ramanathan, and Anne L MartelIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI-W) - Graphs in Biomedical Image Analysis, 2023
Self Supervised Multi-view Graph Representation Learning in Digital PathologyVishwesh Ramanathan, and Anne L MartelIn International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI-W) - Graphs in Biomedical Image Analysis, 2023- Best Paper Award at the MICCAI Workshop on Graphs in Biomedical Image Analysis 2023, held in conjunction with MICCAI 2023 in Vancouver, Canada
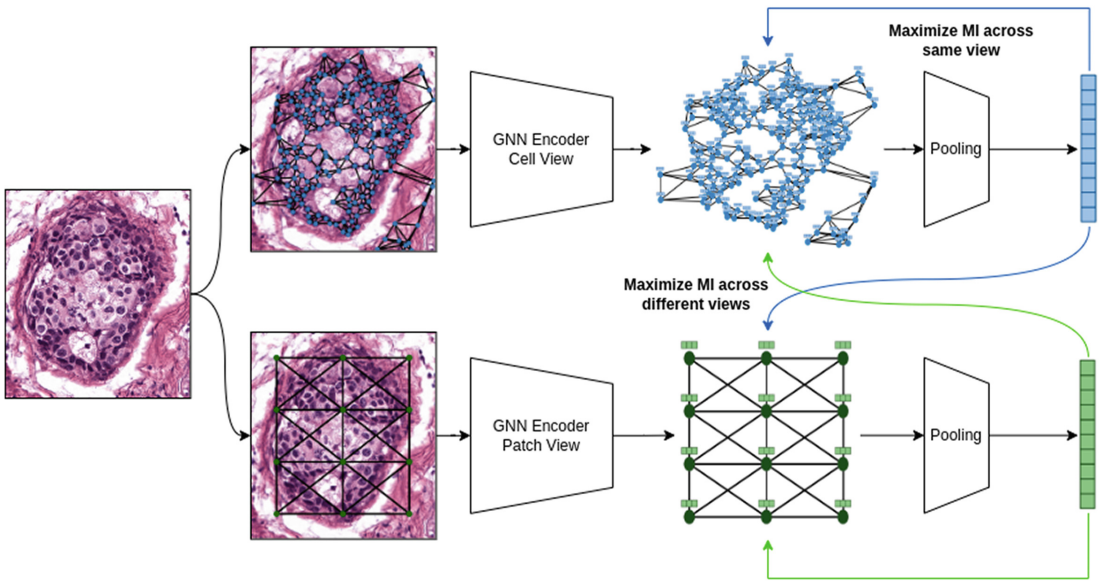
Graph Neural Networks (GNNs) hold great promise for solving many challenges in digital pathology by leveraging the rich relationships between cells and tissues in histology images. However, the shortage of annotated data in digital pathology presents a significant challenge for training GNNs. To address this, self-supervision can be used to enable models to learn from data by capturing rich structures and relationships without requiring annotations. Inspired by pathologists who take multiple views of a histology slide under a microscope for exhaustive analysis, we propose a novel methodology for graph representation learning using self-supervision. Our methodology leverages multiple graph views constructed from a given histology image to capture diverse information. We maximize mutual information across nodes and graph representations of different graph views, resulting in a comprehensive graph representation. We showcase the efficacy of our methodology on the BRACS dataset where our algorithm generates superior representations compared to other self-supervised graph representation learning algorithms and comes close to pathologists and supervised learning algorithms